sfaira - data and model repository for single-cell data¶
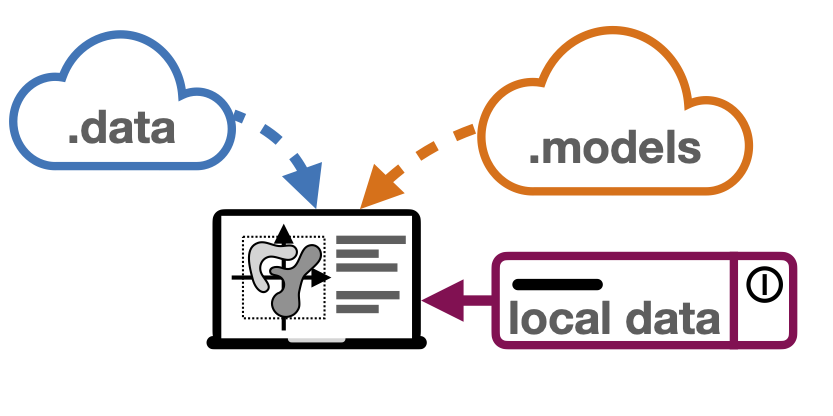
sfaira is a model and a data repository in a single python package. We provide an interactive overview of the current state of the zoos on sfaira-portal.
sfaira fits into an environment of many other project centred on making data and models accessible.
Data zoo¶
We focus on providing a python interface to interact with locally stored data set collections without requiring dedicated data reading and annotation harmonisation scripts: These code blocks are absorbed into our data zoo backend and can be conveniently triggered with short commands.
Model zoo¶
A large body of recent research has been devoted to improving models that learn representation of cell captured with single-cell RNA-seq. These models include embedding models such as autoencoders and cell type prediction models. Many of these models are implemented in software packages and can be deployed on new data sets. In many of these cases, it also makes sense to use pre-trained models to leverage previously published modelling results. We provide a single interface to interact with such pre-trained models which abstracts model settings into a API so that users can easily switch between different pre-trained models. Importantly, model execution is performed locally so that data does not have to be uploaded to external servers and model storage is decentral so that anybody can contribute models easily. Users benefit from easy, streamlined access to models that can be used in analysis workflows, developers benefit from being able to deploy models to a large community of users without having to set up a model zoo.
News¶
No news yet, stay tuned!

